Bayesian Networks
- Probability Overview
- Probability Inference
- Bayesian Network Representation
- Three Canonical Triples
- D-Separation
- Exact Inference
- Approximate Inference
Probability Overview
A random variable represents an event whose outcome is unknown. A probability distribution $P(\omega)$ is assignment of weights to outcomes. A joint distribution is denoted as $P(A, B, C)$. This can be expanded using a chain rule (product rule)
\[P(A_1, A_2, \dots, A_k) = P(A_1)P(A_2 \mid A_1) \dots P(A_k \mid A_1, \dots, A_{k-1})\]The marginal distribution (given a set of $A, B, C$) of $A, B$ can be obtained by summing out all possible values that variable $C$ can take $P(A, B) = \sum_c P(A, B, C = c)$. Similarly, the marginal distribution of $A$ can be obtained as $\sum_b \sum_c P(A, B = b, C = c)$. Conditional probabilities assign probabilities to events conditioned on some known facts $P(A \mid B = b)$.
\[P(A \mid B) = \frac{P(A, B)}{P(B)}\]Combining conditional probability and the chain rule returns the Bayes’ Rule:
\[P(A \mid B) = \frac{P(B \mid A)P(A)}{P(B)}\]$A \perp B$ means these random variables are mutually independent. When $A \perp B$, $P(A, B) = P(A)P(B)$. To write that random variables $A, B$ are conditionally independent given another random variable $C$, we write $A \perp B \mid C$. If $A \perp B \mid C$, then $P(A, B \mid C) = P(A \mid C)P(B \mid C)$. That is, if we have a knowledge of the value of $C$, then $B$ and $A$ do not affect each other. Equivalent definitions include:
\[A \perp B \mid C\] \[P(A \mid B, C) = P(A \mid C)\] \[P(B \mid A, C) = P(B \mid C)\]Probability Inference
It is useful to model the relationship between various nondeterministic events often through a process called probabilitic inference. We assume a model where each possible state for the world has its own probability. For example $P(\text{winter}, 35, \text{cloudy}) = 0.023$. Our model is a joint distribution, a table (e.g., season $S$, temperature $T$, weather $W$) of probabilities that captures the likelihood of each possible outcome also known as an assignment of variables.
Given a joint Probability Density Function (PDF) we can trivially compute any desired probability distribution $P(Q_1 \dots Q_m \mid e_1 \dots e_n)$ using a simple procedure known as inference by enumeration (IBE). We define three types of variables; (1) Query variables $Q_i$ which are unknown and appear on the left side of the conditional, (2) Evidence variables $e_i$ which are observed, have known values, and appear on the right side of the conditional, and (3) Hidden variables which are values present in the overall joint distribution but no in the desired distribution. To do inference by enumeration; we collect all rows consistent with observed evidence variables, sum out (marginalise) all hidden variables, and normalise the table so that is is a probability distribution (summing to 1).
Bayesian Network Representation
Representing an entire joint distribution in memory of a computer is impractical for real problems (exponential space complexity). Bayesian Networks avoid this issue by distributing probabilities across a number of smaller conditional probability tables along with a directed acyclic graph (DAG) to capture the relationship between variables. Formally, the Bayesian Networks consist of a DAG with one node per variable $X$, and a conditional distribution for each node $P(X \mid A_1, \dots, A_n)$, where $A_i$ is the $i$-th parent of $X$, stored as a conditional probability table (CPT). Each CPT has $n+2$ columns consisting of; the value of each of the $n$ parent variables $A_1, \dots, A_n$, the values of $X$, and the conditional probability of $X$ given its parents.
The structure of the Bayesian Network graph encodes conditional independence relations between different nodes. These conditional independences allow us to store multiple small tables instead of one large one. Note that edges in network do not suggest a causal relationship (or dependence) between those nodes, they simple mean there may be some relationship between the nodes. Consider a network representation of five random variables $A, B, E, J, M$. In this Bayesian Network, we would store the following probability tables:
\[P(B), P(E), P(A \mid B, E), P(J \mid A), P(M \mid A)\]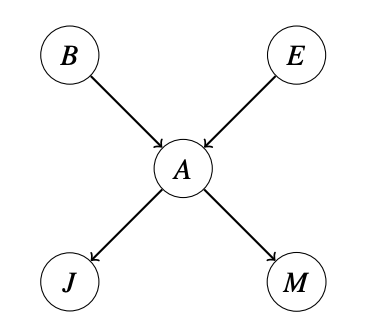
Given all of the CPTs for a graph, we can calculate the probability of a given assignment as follows (and for the above example):
\[P(X_1, X_2, \dots, X_n) = \prod^{n}_{i=1} P(X_i \mid parents(X_i))\] \[P(-b, -e, +a, +j, -m) = P(-b) \cdot P(-e) \cdot P(+a \mid -b, -e) \cdot P(+j \mid +a) \cdot P(-m \mid +a)\]Structure of Bayesian Networks
There are two key rules for Bayesian Network independences:
- Each node is conditionally independent of all its ancestor nodes (or non-descendants) in the graph, given all of its parents.
- Each node is conditionally independent of all other variables given its Markov blanket. A Markov blanket consists of parents, children, and children’s other parents.
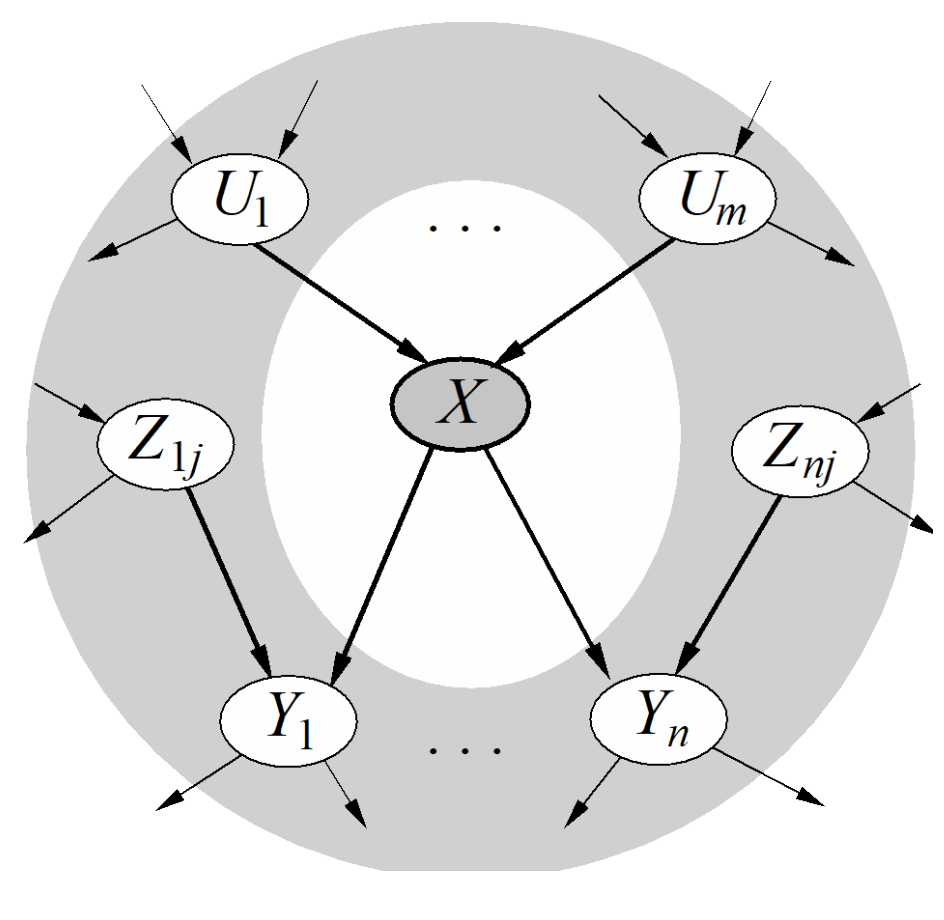
The relation between the joint distribution and the CPTs of the Bayesian Network works because of the conditional independence relationships given by these two rules.
Three Canonical Triples
There are three important canonical cases of connected three-node two-edge Bayesian Networks (aka triples), and the conditional independence relationships they express. A causal chain ($X \rightarrow Y \rightarrow Z$) expresses the following representation of the joint distribution over $X, Y, Z$; $P(x, y, z) = P(z \mid y)P(y \mid x) P(x)$. It is important to note that $X$ and $Z$ are not guaranteed to be independent.
\[P(y \mid x) = \begin{cases} 1 \ \ if \ x = y \\ 0 \ \ else \end{cases}\] \[P(z \mid y) = \begin{cases} 1 \ \ if \ z = y \\ 0 \ \ else \end{cases}\]In this case, $P(z \mid x) = 1$ if $x = z$ and $0$ otherwise, hence $X$ and $Z$ are not independent. To summarise, in the causal chain configuration we can make the statement $X \perp Z \mid Y$.
Another possible configuration for a triple is the common cause ($X \leftarrow Y \rightarrow Z$). It expresses the representation $P(x, y, z) = P(x \mid y)P(z \mid y)P(y)$ (where $y$ is the common cause for $x, z$). Just like the causal chain we can show that $X$ is not guaranteed to be independent of $Z$.
\[P(x \mid y) = \begin{cases} 1 \ \ if \ x = y \\ 0 \ \ else \end{cases}\] \[P(z \mid y) = \begin{cases} 1 \ \ if \ z = y \\ 0 \ \ else \end{cases}\]Then $P(x \mid z) = 1$ if $x = z$ nad otherwise $0$, so $X$ and $Z$ are not independent. And similarly to the causal chain $X \perp Z \mid Y$.
\[P(X \mid Z, y) = \frac{P(X, Z, y)}{P(Z, y)} = \frac{P(X \mid y)P(Z \mid y)P(y)}{P(Z \mid y)P(y)} = P(X \mid y)\]Finally, common effect ($X \rightarrow Y \leftarrow Z$) expresses the representation $P(x, y, z) = P(y \mid x, z)P(x)P(z)$. Again, $X$ and $Y$ are independent $X \perp Z$, however, they are not necessarily independent when conditioned on $Y$. When $Y$ is observed, then knowing $X$ will tell us the value of $Z$, and vice-versa. Common Effect can be viewed as opposite to Causal Chains and Common Cause. $X$ and $Z$ are guaranteed to be independent if $Y$ is not conditioned on. But when conditioned on $Y$, $X$ and $Z$ may be dependent on the specific probability values for $P(Y \mid X, Z)$. The same logic applies when conditioning on descendants of $Y$ in the graph.
D-Separation
We can use the previous three cases as building blocks to help us answer conditional independence questions on an arbitrary Bayesian Network with more than three nodes and two edges. Given a Bayesian Network $G$, two nodes $X$ and $Y$ and a (possibly empty) set of nodes $\set{Z_1, \dots, Z_k}$ that represent observed variables, the following statement must be true $X \perp Y \mid \set{Z_1, \dots, Z_k}$ if these set of nodes d-separate $X$ and $Y$.
D-separation (directed separation) is a property of the structure of the Bayesian Networks that implies this conditional independence relationship, and generalises the cases we have seen in triples. We say that if a set of variables $\set{Z_1, \dots, Z_k}$ d-separates $X$ and $Y$, then $X \perp Y \mid \set{Z_1, \dots, Z_k}$ in all possible distributions that can be encoded by the Bayesian Network. Based on the notion of reachability from node $X$ to node $Y$ we can define the d-separation algorithm:
- Shade all observed nodes $\set{Z_1, \dots, Z_k}$ in the graph
- Enumerate all undirected paths from $X$ to $Y$
- For each path: (a) decompose the path into triples, (b) if all triples are active, this path is active and d-connects $X$ to $Y$
- If no path d-connects (some triples within are passive) $X$ and $Y$, then $X \perp Y \mid \set{Z_1, \dots, Z_k}$.
If all triples are active, then the path is active and d-connects $X$ to $Y$, meaning $X$ is not guaranteed to be conditionally independent of $Y$ given the observed nodes. A triple is active when:
- Casual chain $A \rightarrow B \rightarrow C$ where $B$ is unobserved
- Common cause $A \leftarrow B \rightarrow C$ where $B$ is unobserved
- Common effect $A \rightarrow B \leftarrow C$ where $B$ or one of its descendants is observed
Exact Inference in Bayesian Networks
Inference is the problem of finding the value of some probability distribution $P(Q_1, \dots Q_k \mid e_1 \dots e_k)$. Given a Bayesian Network we can solve this problem naively by forming a joint PDF and using Inference by Enumeration (requiring the creation of and iteration over an exponentially large table).
An alternative approach is to use variable elimination, and eliminate hidden variables one by one. To eliminate a variable $X$ we join all factors involving $X$ and sum out $X$. A factor is defined simply as an unnormalised probability.
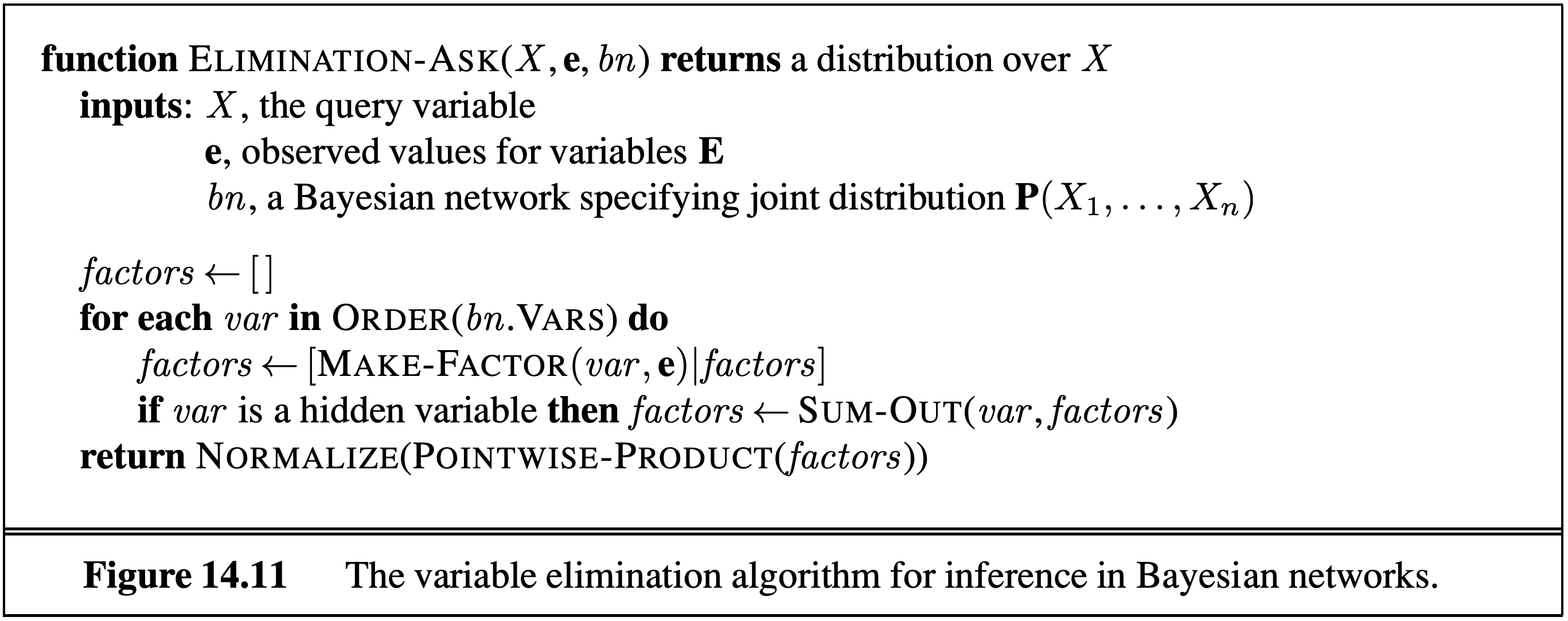

An alternative way of looking at the problem is to observe that (e.g., in set $\set{T, C, S, E}$) the calculation of $P(T \mid +e)$ can either be done through inference by enumeration or by variable elimination as follows:
\[\alpha \sum_s \sum_c P(T) P(s \mid T)P(c \mid T)P(+e \mid, c, s)\] \[\alpha P(T) \sum_s P(s \mid T) \sum_c P(c \mid T) P(+e \mid, c, s)\]Approximate Inference
An alternate approach for probabilistic reasoning is to implicitly calculate the probabilities for our query by counting samples. Suppose we want to calculate $P(+t \mid +e)$. We could collect all samples for which $E = +e$ and compute the fraction of those samples for which $T = +t$, and therefore compute the inference.
Given a Bayesian Network model, we can write a simple simulator that generates $n$ samples from our CPTs also known as prior sampling. The downside of this approach is that it may require the generation of very large number of samples in order to perform analysis of unlikely scenarios. E.g., say $P(+t) = 0.99, \ P(-t)=0.01$ we wanted to compute $P(C \mid -t)$ we would have to throw away 99% of our samples.
One way to mitigate this problem is to modify our procedure to early reject any sample inconsistent with our evidence. For example, for the query $P(C \mid -t)$ we would avoid generating a value for $C$ unless $t$ is false. This means we would have to throw away most of our samples, but at least the bad samples we generate take less time to create. We call this approach rejection sampling.
Likelihood Weighting
A more exotic approach likelihood weighting ensures that we never generate a bad sample. It solves the issue by using a weight for each sample, which is the probability of the evidence variables given the sampled variables. That is, instead of counting all samples equally, we can define a weight $w_j$ for sample $j$ that reflects how likely the observed values for the evidence variables are, given the sampled values. This way we ensure that every CPT participates. To do this, we iterate through each variable in the Bayesian Network (same as normal sampling), sampling a value if the variable is not an evidence variable, or changing the weight for the sample if the variable is evidence.
For the $T, C, S, E$ example, suppose we want to calculate $P(T \mid +c, +e)$. For the $j$-th sample we would perform the following algorithm:
- Set $w_j$ = 1.0, $c = true$, $e = true$
- For $T$ (not evidence) we sample $t_j$ from $P(T)$
- For $C$ (evidence) we update the weigth $w_j = w_j \cdot P(+c \mid t_j)$
- For $S$ we sample $s_j$ from $P(S \mid t_j)$
- For $E$ we update the weight $w_j = w_j \cdot P(+e \mid +c, s_j)$
Then when we perform the usual counting process, we weight sample $j$ by $w_j$ where $0 \leq w_j \leq 1$. This works because in the final calculations for the probabilities, the weights effectively serve to replace the missing CPTs. In effect, we ensure the weighted probability of each sample is given by:
\[P(z_1, \dots, z_p, e_1, \dots, e_m) = \bigg[ \prod^p_{i=1} P(z_i \mid Parents(z_i)) \bigg] \cdot \bigg[ \prod^m_{i=1} P(e_i \mid Parents(e_i)) \bigg]\]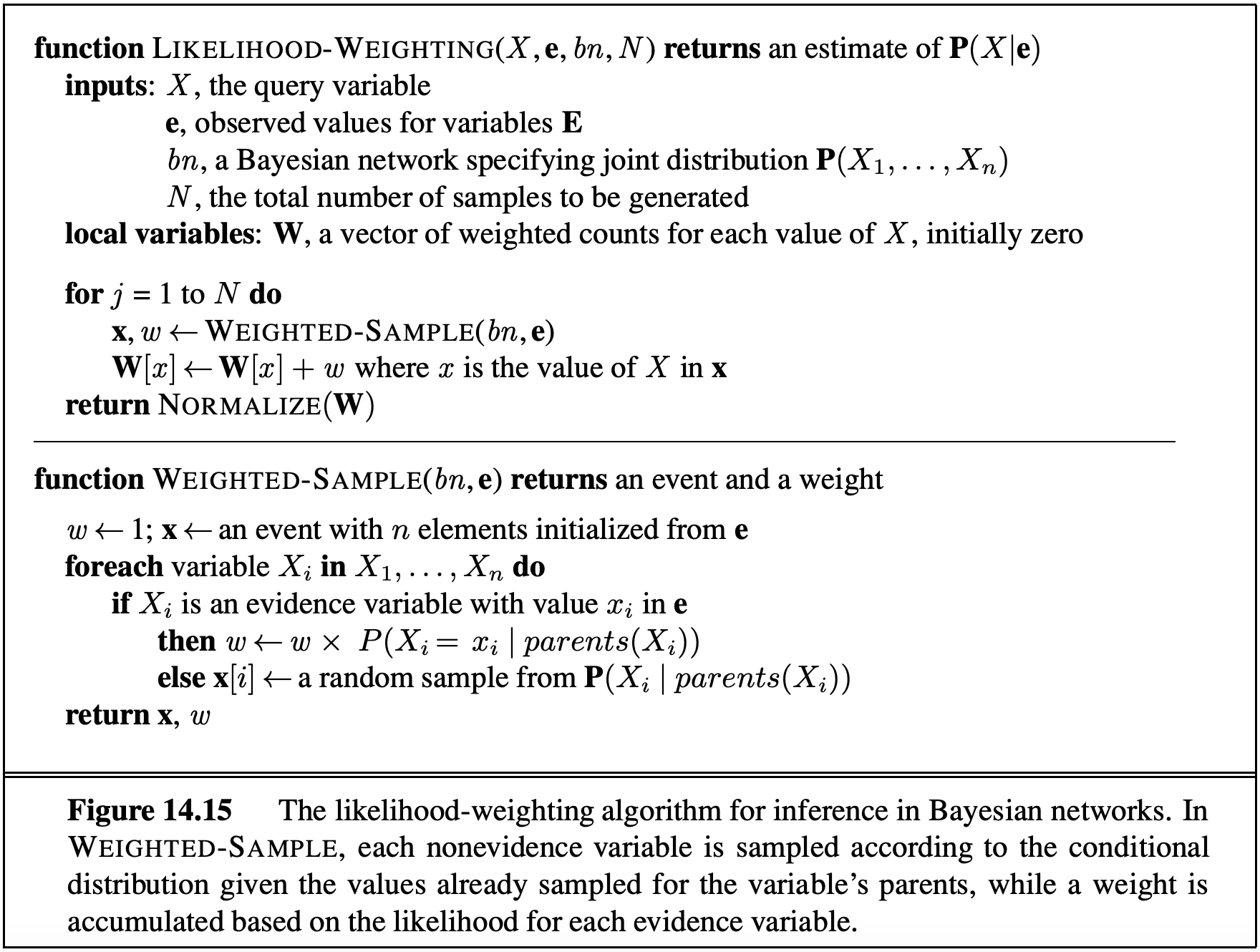
Gibbs Sampling
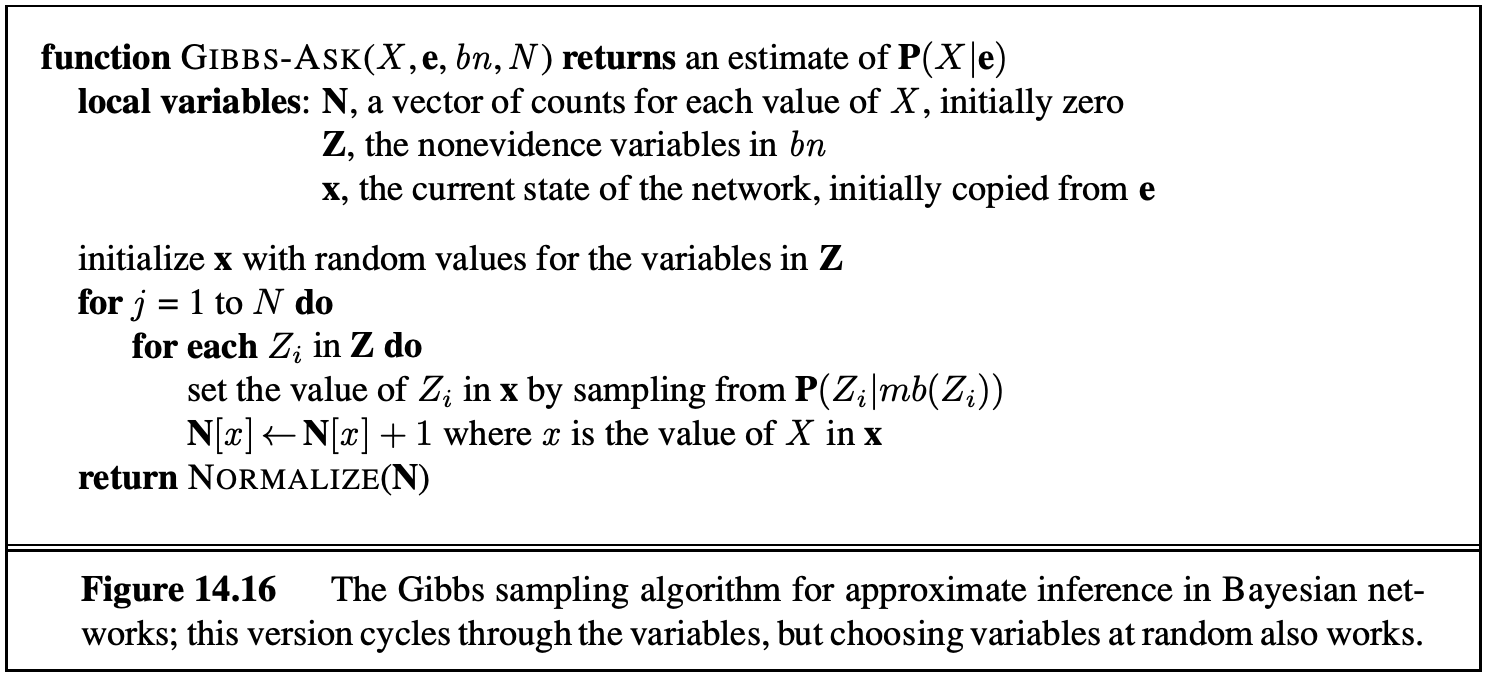
Gibbs Sampling is the fourth approach for sampling. In this approach, we first set all variables to some totally random value. We then repeatedly pick one variable at a time, clear its value, and resample it given the values currently assigned to all other variables. For the $T, C, S, E$ example, we can assign random values $t = true, c = true, s = false, e = true$. We can then pick one of our four variables to resample say $S$. Using our distribution (randomly initalised) we pick a new variable $P(S \mid +t, +c, +e)$. This requires the knowledge of $P(S \mid T, C, E)$ which can be calculated using only the CPTs that connect $S$ with its neighbours. Thus, in networks with small number of neighbours we can precompute conditional distributions (for each variable) in linear time. With enough repetition our later samples converge to the correct distribution.
 </img>
</img>

Summary
- Bayesian Networks is a powerful representation of joint probability distributions, and can be used to model arbitrary distributions to perform inference and sampling.
- Inference By Enumation and Variable Elimination also known as exact inference, guarantee exact correct probability but are computationally expensive.
- Sampling methods (Prior, Rejection, Likelihood, Gibbs) are used too approximate solutions while using less compute.
References
Enjoy Reading This Article?
Here are some more articles you might like to read next: